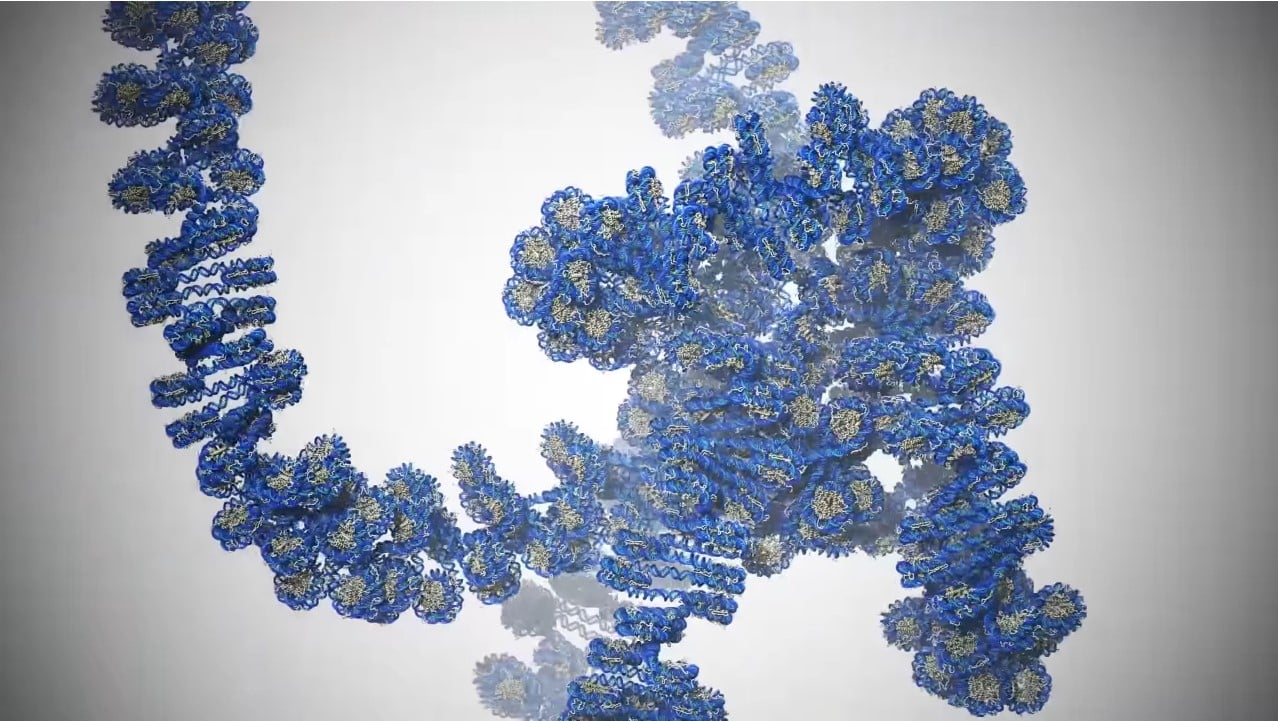
Researchers at Los Alamos National Laboratory have simulated all the atoms within a gene in DNA, which amounts to 1 billion atoms. This represents the largest and most accurate simulation to date, and it could help researchers on the hunt for cures for diseases such as cancer.
“It is important to understand DNA at this level of detail because we want to understand precisely how genes turn on and off,” Los Alamos structural biologist Karissa Sanbonmatsu said in a statement. “Knowing how this happens could unlock the secrets to how many diseases occur.”
In this experiment, researchers used all the atoms within a gene in DNA to learn more about how DNA expands and contracts, controlling the on/off switches in human genes.
Sanbonmatsu and her colleagues modeled and ran the model on Los Alamos’ Trinity supercomputer, which is ranked as the sixth fastest in the world. Its capabilities support the National Security Administration’s stockpile stewardship program, which ensures these experiments are safe, secure and effective.
DNA is an essential material in all living things on Earth with genes that show body structures and activities. According to the researchers at Los Alamos, there’s enough DNA in a single human body to wrap our planet 2.5 million times.
A long, string-like DNA molecule is joined in a network of small molecular spools, and their winding and unwinding turns genes on and off. Researchers want to learn more about this spool network, which is also called epigenetics, a new field of science which studies the development of bodies in the womb and how diseases form.
Simulating all the atoms within a gene in DNA helps researchers solve the mystery of how genes spread and contract, but it’s an extremely challenging task which requires a lot of computational resources.
“Right now, we were able to model an entire gene with the help of the Trinity supercomputer at Los Alamos,” Los Alamos polymer physicist Anna Lappala Los Alamos said. “In the future, we’ll be able to make use of exascale supercomputers, which will give us a chance to model the full genome.”
Researchers also highlight the importance of exascale computers as the next generation of supercomputers, which are expected to run calculations much faster than currently-available machines. Future supercomputers should enable researchers to model an entire human genome, which should shed more light on how genes are turned on and off.
The study describing these experiments was published in the Journal of Computational Chemistry on Apr. 17. In addition to the Los Alamos team, researchers from the RIKEN Center for Computational Science in Japan, the New Mexico Consortium and New York University also worked on it.