MIT Researchers collaborated with the Arizona State University to create a computer program which translates 2D drawing into DNA structure. The program is capable of translating any free-form 2D drawing into a two-dimensional nanoscale structure which consists of DNA.
The team used a new program which allows anyone to create DNA nanostructures of any shape, used in cell biology, photonics and quantum sensing and computing and more.
“What this work does is allow anyone to draw literally any 2-D shape and convert it into DNA origami automatically,” says Mark Bathe, an associate professor of biological engineering at MIT and the senior author of the study said in a news release, describing the findings published on Jan. 2 in Science Advances. The program is called PERDIX and can be found online. Other lead authors of the paper include Hyungmin Jun, a MIT postdoc, and Fei Zhang, an assistant research professor at Arizona State University.
The idea of DNA origami originates from the early 1980’s, and is a discipline including folding DNA into small structures. There were many approaches that came later and included creating complex 3D DNA structures, although all the structures were more than challenging to make. In 2016, Bathe and his team created a way which allowed them to automate making 3D polyhedral DNA structures. The new study aims to automate how 2D DNA structures work.
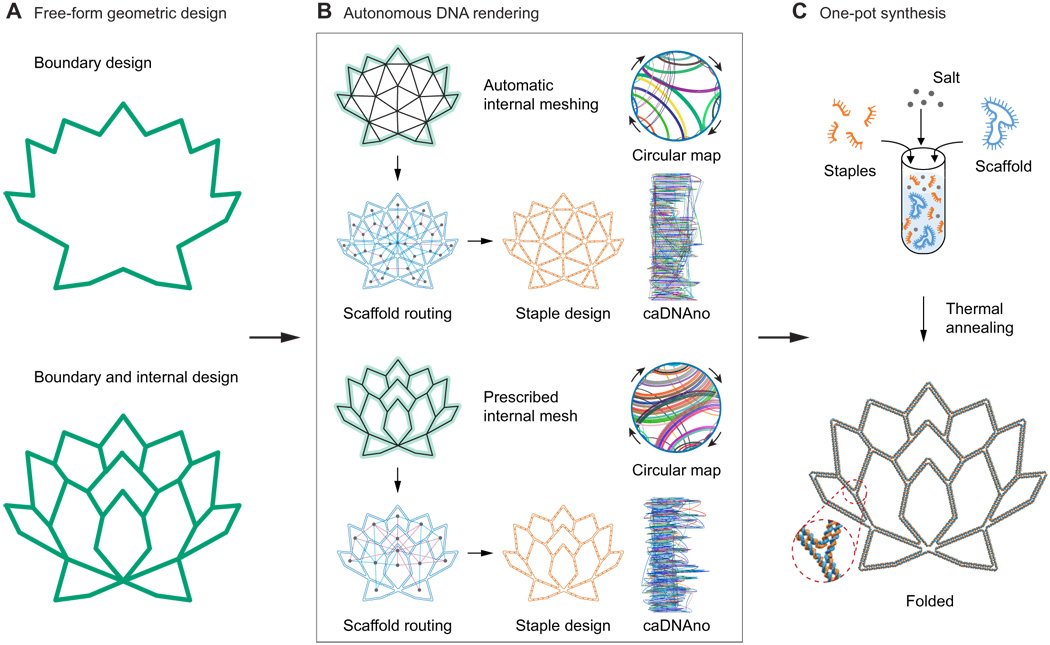
They used a new mathematical model which can manage the process of routing the single-stranded scaffold, making the entire structure create a correct shape. With proper instructions, the program translates 2D drawing into DNA structure, of any free form.
The free-form 2D shape can be created on any computer drawing program and converted into a computer-aided design (CAD) file. Then, that drawing gets fed into the DNA design program.
“Once you have that file, everything’s automatic, much like printing, but here the ink is DNA,” Bathe said.
When the drawings are sequenced, users can order them to resemble some specific shape. The researchers created shapes which have edges consisting of two duplexes of DNA. However, the team also has a program which can create six duplexes per edge, earning more rigidness. There is also software for 3D polyhedra called TALOS, which is available online and will be added in the journal ACS Nano.
“The fact that we can design and fabricate these in a very simple way helps to solve a major bottleneck in our field,” Bathe said. “Now the field can transition toward much broader groups of people in industry and academia being able to functionalize DNA structures and deploy them for diverse applications.”